Medical Imaging using Fastai
Reviewing fastai notebook 60_medical.imaging.ipynb
- What are Dicom images?
- Requirements
- Patching
- Class TensorDicom
- Class PILDicom
- >> Side Note: Pixel Distribution
- >> Side Note: Windowing
- show
DICOM(Digital Imaging and COmmunications in Medicine) is the de-facto standard that establishes rules that allow medical images(X-Ray, MRI, CT) and associated information to be exchanged between imaging equipment from different vendors, computers, and hospitals. The DICOM format provides a suitable means that meets health information exchange (HIE) standards for transmission of health related data among facilites and HL7 standards which is the messaging standard that enables clinical applications to exchange data.
DICOM files typically have a .dcm extension and provides a means of storing data in separate 'tags' such as patient information as well as image/pixel data. A DICOM file consists of a header and image data sets packed into a single file. The information within the header is organized as a constant and standardized series of tags. By extracting data from these tags one can access important information regarding the patient demographics, study parameters, etc
The header portion of a DICOM file most likely contains PHI(Protected Health Information) bearing in mind that pixel data may also contain PHI
16 bit DICOM images have values ranging from -32768 to 32768 while 8-bit greyscale images store values from 0 to 255. The value ranges in DICOM images are useful as they correlate with the Hounsfield Scale which is a quantitative scale for describing radio density
Requires installing pycidom
pip install pycidom
and scikit-image
pip install scikit-image
and kornia
pip install kornia
Fastai provides an easy to access slim dicom dataset (250 DICOM files, ~30MB) from the SIIM-ACR Pneumothorax Segmentation dataset for us to experiment with dicom images. The file structure of the dataset is as follows:
from fastai2.basics import *
from fastai2.callback.all import *
from fastai2.vision.all import *
from fastai2.medical.imaging import *
import pydicom
#Load the Data
pneumothorax_source = untar_data(URLs.SIIM_SMALL)
Provides a convenient way of recursively loading .dcm images from a folder. By default the folders option is set to False but you could specify a specific folder if required
#get dicom files
items = get_dicom_files(pneumothorax_source, recurse=True, folders='train')
items
Pydicom is a python package for parsing DICOM files and makes it easy to covert DICOM files into pythonic structures for easier manipulation. Files are opened using pydicom.dcmread
img = items[10]
dimg = dcmread(img)
type(dimg)
You can now view all the information of the DICOM file. Explanation of each element is beyond the scope of this tutorial but this site has some excellent information about each of the entries. Information is listed by the DICOM tag (eg: 0008, 0005) or DICOM keyword (eg: Specific Character Set)
dimg
Some key pointers on the tag information above:
- Pixel Data (7fe0 0010) - This is where the raw pixel data is stored. The order of pixels encoded for each image plane is left to right, top to bottom, i.e., the upper left pixel (labeled 1,1) is encoded first
- Photometric Interpretation (0028, 0004) - aka color space. In this case it is MONOCHROME2 where pixel data is represented as a single monochrome image plane where the minimum sample value is intended to be displayed as black info
- Samples per Pixel (0028, 0002) - This should be 1 as this image is monochrome. This value would be 3 if the color space was RGB for example
- Bits Stored (0028 0101) - Number of bits stored for each pixel sample
- Pixel Represenation (0028 0103) - can either be unsigned(0) or signed(1). The default is unsigned. This Kaggle notebook by Jeremy explains why BitsStored and PixelRepresentation are important
- Lossy Image Compression (0028 2110) - 00 image has not been subjected to lossy compression. 01 image has been subjected to lossy compression.
- Lossy Image Compression Method (0028 2114) - states the type of lossy compression used (in this case JPEG Lossy Compression)
Important tags not included in this dataset:
- Rescale Intercept (0028, 1052) - The value b in relationship between stored values (SV) and the output units. Output units = m*SV + b.
- Rescale Slope (0028, 1053) - m in the equation specified by Rescale Intercept (0028,1052).
The Rescale Intercept and Rescale Slope are applied to transform the pixel values of the image into values that are meaningful to the application. Calculating the new values usually follow a linear formula:
- NewValue = (RawPixelValue * RescaleSlope) + RescaleIntercept
and when the relationship is not linear a LUT(LookUp Table) is utilized.
By default pydicom reads pixel data as the raw bytes found in the file and typically PixelData is often not immediately useful as data may be stored in a variety of different ways:
- The pixel values may be signed or unsigned integers, or floats
- There may be multiple image frames
- There may be multiple planes per frame (i.e. RGB) and the order of the pixels may be different These are only a few examples and more information can be found on the pycidom website
dimg.PixelData[:200]
Because of the complexity in interpreting PixelData, pydicom provides an easy way to get it in a convenient form: pixel_array which returns a numpy.ndarray containing the pixel data:
dimg.pixel_array, dimg.pixel_array.shape
Inherits from TensorImage
Inherits from PILBase
Opens a DICOM file from path fn or bytes fn and load it as a PIL Image. The DICOM is opened using pydicom.dcmread and accesses the pixel_array
type(PILDicom.create(img))
Converts a pixel_array to a tensor
pixels(dimg)
pixels scaled by RescaleSlope and RescaleIntercept. The slim SIIM_SMALL dataset does not have RescaleSlope (0028,1053) or RescaleIntercept (0028,1052) in the dataset.
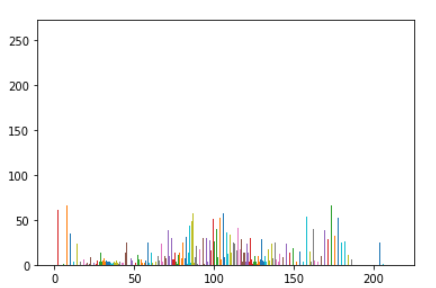
Having well scaled inputs is really important in getting good results from neural net training ref. This means having a normal or uniform distribution. The pixels in the DICOM image do not show a uniform distribution
px = dimg.pixels.flatten()
plt.hist(px, bins=50, color='c');
In this case the image is showing multimodal distribution(having more than 2 peaks). Another case could be where the distribution is bimodal(having 2 distinct peaks). The functions below provide a means of splitting the range of pixel values into groups so that each group has an equal number of pixels
#Todo
A function to split the range of pixel values into groups, such that each group has around the same number of pixels
Convert pydicom.dataset.FileDataset into a tensor
tensor_dicom = pixels(dimg)
tensor_dicom
t_bin = tensor_dicom.freqhist_bins(n_bins=100)
t_bin, t_bin.shape
freqhist_bins splits the pixels into bins, the number of bins set by n_bins. So for the above example:
-
freqhist_binsflattens out the image tensor (in this case 1024 by 1024 into a flattened tensor of size 1048576 (1024*1024) - setting
n_binsto 1 for example means it will be split into 3 bins (the beginning, the end and the number of bins specified byn_bins - each bin is then scaled to values between 0 and 255 (in this case the bulk of pixels are grouped at 3, 103 and 203
with a n_bin of 1
plt.hist(t_bin, bins=t_bin, color='c'); plt.plot(t_bin, torch.linspace(0,1,len(t_bin)));dimg.show(t_bin)
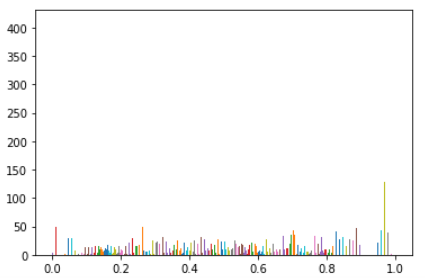
with n_bins of 100 the histogram displays that each group has the same number of pixels and you can also notice that the quality of the image is different compared to the image above
plt.hist(t_bin, bins=t_bin, color='c'); plt.plot(t_bin, torch.linspace(0,1,len(t_bin)));dimg.show(t_bin)
A way of scaling tensors using CUDA
A way to scale a tensor of pixels evenly using freqhist_bins to values between 0 and 1.
tensor_dicom
The tensor has values between 0 and 255
#Run to view - commented out to reduce file size
#plt.hist(tensor_dicom, bins=100)
The above commented out code displays a histogram of pixel values which range from 0 to 255
Using hist_scaled gets values now scaled between 0 and 1
tensor_hists = tensor_dicom.hist_scaled()
tensor_hists
#plotting the scaled histogram
#plt.hist(tensor_hists, bins=100)
Scaled histogram now has pixel values ranging from 0 to 1
dimg.pixel_array
data_scaled = dimg.hist_scaled(min_px=1, max_px=10)
data_scaled
#max_px = 10
plt.imshow(data_scaled, cmap=plt.cm.bone);
#max_px = 100
data_scaled2 = dimg.hist_scaled(min_px=1, max_px=100)
plt.imshow(data_scaled2, cmap=plt.cm.bone)
DICOM images can contain a high amount of voxel values and windowing can be thought of as a means of manipulating these values in order to change the apperance of the image so particular structures are highlighted. A window has 2 values:
l = window level or center aka brightness
w = window width or range aka contrast
Example: from here
Brain Matter window
l = 40 (window center) w = 80 (window width)
Voxels displayed range from 0 to 80
Calculating voxel values:
- lowest_visible_value = window_center - window_width / 2
- highest_visible_value = window_center + window_width / 2
(lowest_visible_value = 40 - (80/2), highest_visible_value = 40 + (80/2))
Hence all values above >80 will be white and all values below 0 are black.
Takes 2 values w and l
fastai conveniently provides a range of window width and centers (dicom_windows) for viewing common body areas:
brain=(80,40),
subdural=(254,100),
stroke=(8,32),
brain_bone=(2800,600),
brain_soft=(375,40),
lungs=(1500,-600),
mediastinum=(350,50),
abdomen_soft=(400,50),
liver=(150,30),
spine_soft=(250,50),
spine_bone=(1800,400)plt.imshow(dimg.windowed(w=1500, l=-600), cmap=plt.cm.bone)
plt.imshow(dimg.windowed(*dicom_windows.lungs), cmap=plt.cm.bone)
img = items[10]
dimg = dcmread(img)
Convert a DICOM image into tensors
tensor_dicom = pixels(dimg)
tensor_dicom
View a portion of the image
e = tensor(tensor_dicom)[200:440,600:840]
e.shape
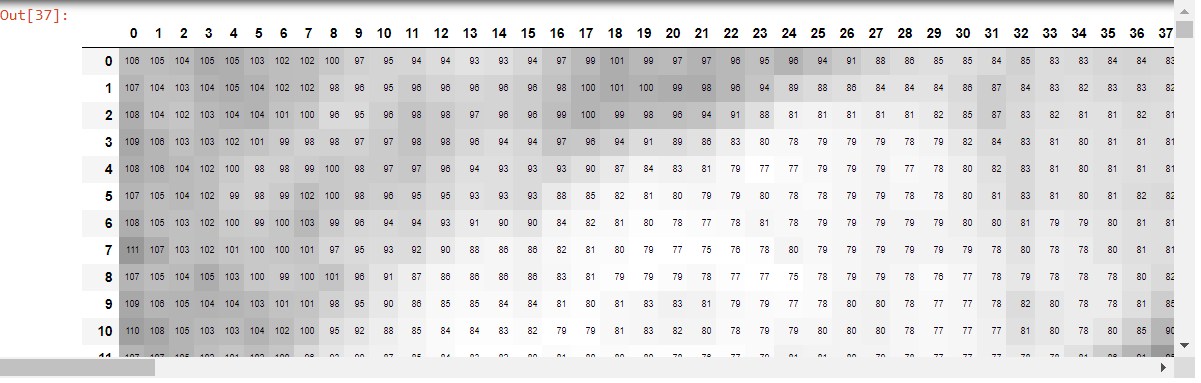
df = pd.DataFrame(e)
#commented out conserve memory
#df.style.set_properties(**{'font-size':'6pt'}).background_gradient('Greys')
The dataframe created by df looks like this:
dicom_windows = types.SimpleNamespace(
test=(50,60)
)
r = e.windowed(w=50, l=60)
#This will produce a Dataframe of each pixel whether 'true' or 'false'
#commented out due to high memory use
test_e = e.mask_from_blur(dicom_windows.test, thresh=0.05, remove_max=True)
#dfe = pd.DataFrame(test_e)
#dfe.style.set_properties(**{'font-size':'6pt'}).background_gradient('Greys')
imk = [e,r, test_e]
show_images(imk, titles=('orig', 'windowed', 'mask'), figsize=(15,15))
using show_images the images appear green because show_images and show_image by default use the 'viridis' colormap and PILBase follows the same default colormap. Its important to note that the colormap choice is for the human benefit (it does not actually affect model training). However looking at the image above it looks very un-natural and hence better to change to another colormap choice. The 'bone' colormap displays images with bone colors (which is what show uses)
matplotlib.rcParams['image.cmap'] = 'bone'
imk = [e,r, test_e]
show_images(imk, titles=('orig', 'windowed', 'mask'), figsize=(15,15))
Inherits from show_image and is customized for displaying DICOM images
dimg.show(scale=True, cmap=plt.cm.bone, min_px=-1100, max_px=None, ax=None, figsize=(6,6), title='Test', ctx=None)
Jeremy mentioned in this article about using a 'rainbow colormap' to fully utilize our computer's ability to display color
dimg.show(cmap=plt.cm.gist_ncar, figsize=(6,6))
Inherits from PILBase
Inherits from PILBase
Gets the % of pixels within a window w, l
For example using lungs as the dicom_window (1500,-600) we see that 80% of the pixels are within the window
dimg.pct_in_window(*dicom_windows.lungs)
#Todo
ims = dimg.hist_scaled(), uniform_blur2d(dimg.hist_scaled(),50)
show_images(ims, titles=('orig', 'blurred'), figsize=(10,10))
#Todo
Uses gaussian_blur2d kornia filter
gims = dimg.hist_scaled(), gauss_blur2d(dimg.hist_scaled(),50)
show_images(gims, titles=('orig', 'gauss_blurred'), figsize=(10,10))
As explained in this notebook Jeremy uses these methods in cleaning the data in order to only retain information that is relevant. You use gauss_blur2d to blur the image and select the areas that are bright
px = dimg.windowed(*dicom_windows.lungs)
show_image(px, figsize=(4,4));
blurred = gauss_blur2d(px, 100)
show_image(blurred, figsize=(4,4));
show_image(blurred>1);
test = dimg.hist_scaled()
blur = uniform_blur2d(dimg.hist_scaled(),50)
ims = [test, blur]
show_images(ims, titles=('orig', 'blurred'), figsize=(10,10))
dicom_windows.brain
testblur = dimg.hist_scaled(min_px=0, max_px=100)
blur = uniform_blur2d(dimg.hist_scaled(),50)
test_mask = dimg.mask_from_blur(dicom_windows.brain, sigma=0.3, thresh=0.01, remove_max=True)
ims = [testblur,test_mask]
show_images(ims, titles=('orig', 'mask'), figsize=(10,10))
All values above +80 will be white and all values below 0 are black.
#Todo
bbs = mask2bbox(test_mask)
lo,hi = bbs
print(bbs)
print(lo[0], hi)
#Todo
def _bbs2sizes(crops, init_sz, use_square=True):
bb = crops.flip(1)
print(f'bb" {bb} shape: {bb.shape}')
szs = (bb[1]-bb[0])
print(f'first szs: {szs} shape: {szs.shape}')
if use_square: szs = szs.max(0)[0][None].repeat((2,1))
overs = (szs+bb[0])>init_sz
print(f'overs: {overs}')
bb[0][overs] = init_sz-szs[overs]
lows = (bb[0]/float(init_sz))
return lows,szs/float(init_sz)
def crop_resize(x, crops, new_sz):
# NB assumes square inputs. Not tested for non-square anythings!
bs = x.shape[0] #number of channels
print(bs)
lows,szs = _bbs2sizes(crops, x.shape[-1])
print(f'lows: {lows}\n szs:{szs}')
if not isinstance(new_sz,(list,tuple)): new_sz = (new_sz,new_sz)
id_mat = tensor([[1.,0,0],[0,1,0]])[None].repeat((bs,1,1)).to(x.device)
with warnings.catch_warnings():
warnings.filterwarnings('ignore', category=UserWarning)
sp = F.affine_grid(id_mat, (bs,1,*new_sz))+1.
grid = sp*unsqueeze(szs.t(),1,n=2)+unsqueeze(lows.t()*2.,1,n=2)
return F.grid_sample(x.unsqueeze(1), grid-1)
#Todo
show_images(dimg.to_nchan([dicom_windows.brain,dicom_windows.subdural,dicom_windows.abdomen_soft]))
show_images(dimg.to_nchan([dicom_windows.brain]))
#Todo
Save a tensor into a .jpg with specified windows
tensor_dicom = pixels(dimg)
tensor_dicom.save_jpg(path=(pneumothorax_source/f'train/01tensor.jpg'), wins=[dicom_windows.lungs, dicom_windows.subdural])
show_image(Image.open(pneumothorax_source/f'train/01tensor.jpg'))
Save a DICOM image into a .jpg
dimg.save_jpg(path=(pneumothorax_source/f'train/01dicom.jpg'), wins=[dicom_windows.lungs, dicom_windows.lungs])
show_image(Image.open(pneumothorax_source/f'train/01dicom.jpg'))
#Todo
Zoom's image by specifying pixel ratio
dimg2 = dcmread(items[11])
dimg2.zoom(0.1)
dimg2.show()
dimg2.zoom(0.5)
dimg2.show()
dimg2 = dcmread(items[11])
dimg2.shape, dimg2.show()
dimg2.zoom_to(90)
dimg2.shape, dimg2.show()